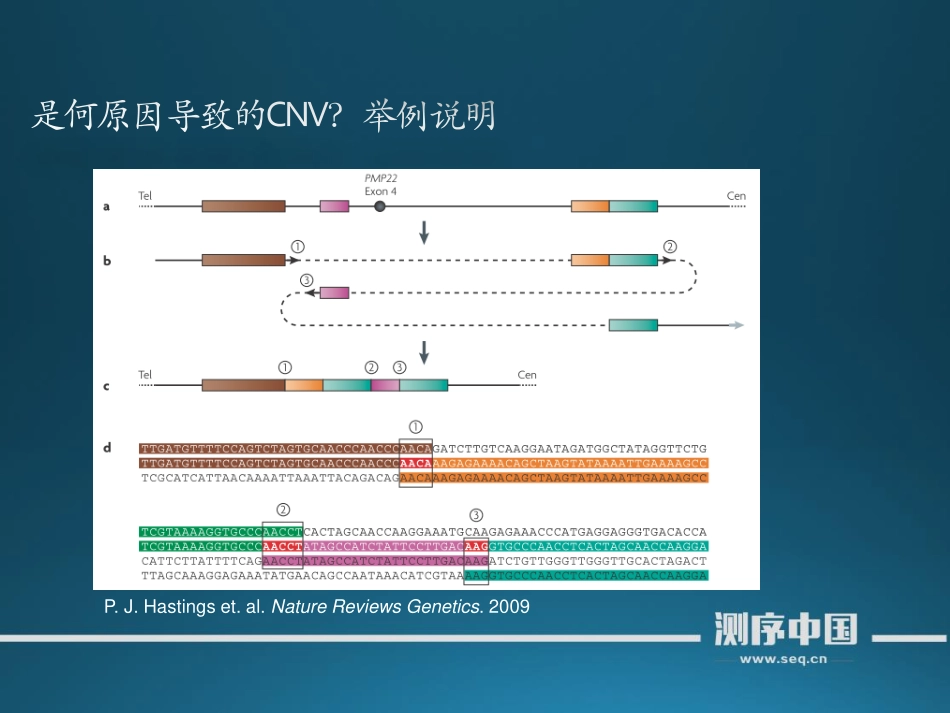
Deletionsandduplicationsofchromosomalsegments(copynumbervariants,CNVs)areamajorsourceofvariationbetweenindividualhumansandareanunderlyingfactorinhumanevolutionandinmanydiseases,includingmentalillness,developmentaldisordersandcancerP.J.Hastingset.al.NatureReviewsGenetics.2009P.J.Hastingset.al.NatureReviewsGenetics.2009芯片NGS$samtoolsview-F4Normal.Bamchr1|perl-lane'print"$F[2]\t$F[3]"'>NormalBam.hit$samtoolsview-F4Tumor.Bamchr1|perl-lane'print"$F[2]\t$F[3]"'>TumorBam.hit$perlcnv-seq.pl--testtumorBam.hit–refnormalBam.hit--genome-size249250621--window-size10000鉴定CNV方法举例:CNV-seq,anewmethodtodetectcopynumbervariationusinghigh-throughputsequencing软件地址:http://tiger.dbs.nus.edu.sg/CNV-seq/$R#inRcommandprompt>library(cnv)data<-read.delim(“somatic.cnv")>cnv.print(data)#output...>cnv.summary(data)#output...>plot.cnv(data,CNV=4,upstream=4e+6,downstream=4e+6)>ggsave("sample.pdf")ExomeCNVhttp://bioinformatics.oxfordjournals.org/content/27/19/2648.longSeqseghttp://www.broadinstitute.org/cgi-bin/cancer/publications/pub_paper.cgi?mode=view&paper_id=182CNVnatorhttp://sv.gersteinlab.org/cnvnator/CNVerhttp://compbio.cs.toronto.edu/cnver/1.Getsoft-clippingpositions*PerlextractSClip.pl-itumor.bam-ref_genomehg19_genome.faPerlextractSClip.pl-inormal.bam-ref_genomehg19_genome.fa2.RemovegermlineeventsPerlcountDiff.pl-dtumor.bam.cover-ggermline.bam.cover3.RunningtheSVdetectionscript.*PerlCREST.pl-fsomatic.cover-dtumor.bam-ggermline.bam–refhg19_genome-thg19_genome.2bitBreakdancerhttp://gmt.genome.wustl.edu/packages/breakdancer/Pindelhttp://gmt.genome.wustl.edu/packages/pindel/谢谢!祝大家2015元旦快乐!微信测序讨论群